Passage in Vero Cells: The Variant Game
“In addition, passage in cell culture can result in artificial mutations in the sequences, which were not present in the original clinical sample. This can have major implications for subsequent analyses. Using cell culture solely for the purpose of amplifying virus genetic material for SARS-CoV-2 sequencing should therefore be avoided“
The above statement comes from a “SARS-COV-2” Genome Sequencing manual from the WHO. They seemingly admit that the cell culture process results in the mutations and variations seen in the over 3 million “SARS-COV-2” variants currently running around at the time of this writing. They claim that this is due to passaging of cell cultures. What exactly is the WHO referring to here?
“Subculturing, also referred to as passaging cells, is the removal of the medium and transfer of cells from a previous culture into fresh growth medium, a procedure that enables the further propagation of the cell line or cell strain.”
This process of transferring cells from one culture to another while changing up the chemicals/medium used is common practice. They do this numerous times in order to keep the “virus” within the cell culture soup “alive” indefinitely. However, this process of removing the cells from one petri dish to another is very stressful on the cells and the addition of fresh cell-altering and DNA-damaging chemicals only heightens this stress. This practice inevitably leads to more damage/cell death and changes the morphology of the cell culture soup the longer this process carries on.

Passage Number Effects in Cell Lines
“Cell lines at high passage numbers experience alterations in morphology, response to stimuli, growth rates, protein expression and transfection efficiency, compared to lower passage cells.
The scientific community is taking notice that cell line quality is crucial to successful experimentation and that avoiding the use of cell lines that have been in culture too long is an important step to ensure reliable and reproducible results. But while the evidence for passage number-related effects on cell lines is compelling, much less is understood about the mechanisms underlying passage-dependent changes and about actions researchers can take to avoid passage number effects in their experiments.”
How many passages are too many?
“A straightforward method for determining the passage number of a cell line does not exist. A review of the literature on passage-related effects in cell lines demonstrates that the effects are complex and heavily dependent on a host of factors such as the type of cell line, the tissue and species of origin, the culture conditions and the application for which the cells are used.”
It is clear that the culture conditions are a major factor in the data generated from the cell culture supernatant. Virologists are essentially creating new sequences/mutations every time they whip up their witches brew. In the case of “SARS-COV-2,” this is highlighted by passages (pun somewhat intended) from two recent studies:

A cautionary perspective regarding the isolation and serial propagation of SARS-CoV-2 in Vero cells
“An array of SARS-CoV-2 virus variants have been isolated, propagated and used in in vitro assays, in vivo animal studies and human clinical trials. Ensuring the genetic stability of SARS-CoV-2 during in vitro propagation is essential but has been too frequently overlooked. Our observations of working stocks of SARS-CoV-2 suggest that sequential propagation in Vero cells leads to critical changes in the region of the furin cleavage site (FCS), which significantly reduce the value of the working stock for critical research studies, vaccine development, production, evaluation and use.”
“The authors of this paper, members of the WHO working group on SARS-CoV-2 virus propagation in cell lines, have pooled the results of carefully analysed genetic data generated from sequencing multiple isolates of serially propagated SARS-CoV-2 in different cell types. Serially propagating SARS-CoV-2 in Vero E6 cells leads to rapid increases in genetic variants, particularly those located in the sequence coding for the FCS of the spike protein.”
Early findings
“In the early phase of the global response to SARS-CoV-2, quality assurance measures taken by Public Health England (PHE) to check the England 02 isolate provided to the Biodefense and Emerging Infections Resources (BEI Resources) included deep sequencing of the second Vero E6 cell line passage of this isolate. This analysis indicated that, although the first passage (P1) stock had no detectable changes (<1%, EPI_ISL_40703) (Table 1), over 90% of the virus content of the P2 stock and 100% of the P3 stock (multiplicity of infection (MOI) ranging from 1.0E−02 to 1.0E−03) contained a 24-nucleotide in-frame deletion in the spike region resulting in loss of 8 amino acids including the FCS (see details in Supplementary Table 1). This observation raised concern among virologists that SARS-CoV-2 isolates being propagated and studied around the globe were not accurately representing the virus circulating in humans.”
“In addition, we noted that a P2 stock propagated in Calu-1 cells did not lose these variants when grown in Vero/SLAM cells but seemed to retain low levels during propagation, whereas the levels of FCS variants rose rapidly in Vero E6 cells irrespective of their source (Table 1).”
Discussion
“Studies conducted at PHE, NIBSC, University of Wisconsin-Madison and BEI Resources all conclude that, when SARS-CoV-2 is propagated in Vero E6 cells, there is a risk that during the sequential passage of this virus for working stock generation, deletions may arise in critical virulence components of the virus, including the FCS. Such deletions appear to result in the stock virus being less virulent in animal models (as measured by clinical observations and/or viral titration in mucosal secretions).”
“On the basis of this preliminary data, we encourage researchers producing stocks of SARS-CoV-2 to consider:
limitation of the number of passages in cell culture, using low MOI, in an effort to maintain wild-type properties;
evaluation and selection of a cell line that supports viral isolation and working stock production with acceptable (<1%) variant thresholds for downstream use;
evaluation of both the consensus sequence and inclusion of analysis of minor variants of each virus preparation;
incorporation of LoFreq4 (or equivalent) sequencing analysis for low-frequency variant calling.”
“Spontaneous mutations due to virus adaptation to both Vero and Vero E6 cells have also been reported for viruses such as Ebola virus and Zika virus.8,9 However, deletions and mutations in the SARS-CoV-2 FCS became so frequently observed in passages 4 and 5 that they dominated the reads taken from workings stocks by up to 99% (Tables 1 and 3). The results at passage 4 were, however, variable such that the FCS region of two different passage 4 stock contained mutations at a frequency of ≈16% in one stock but <1% for another (Table 3). This latter stock when taken to passage 5 did, however, yield a stock with >10% FCS variants. These data suggest that even the same passage level of virus can exhibit entirely different genetic characteristics, further emphasizing that investigators need to confirm the genetic sequence after propagation rather than relying on the sequence of the seed stock.
The findings of this group in this publication support the observations of other groups2,10,11 that FCS changes occur during serial propagation of SARS-CoV-2 in some cell lines. Despite the publication of these articles, there is continued production and dissemination of stocks of virus that are compromised in this manner, especially as there is a perceived need to rapidly isolate and distribute new variants with a combination of changes in the spike protein.”
https://www.nature.com/articles/s41541-021-00346-z
In Summary (Part 1):
- Ensuring the genetic stability of “SARS-CoV-2” during in vitro propagation is essential but has been too frequently overlooked
- Sequential propagation in Vero cells leads to critical changes in the region of the furin cleavage site (FCS)
- Serially propagating “SARS-CoV-2” in Vero E6 cells leads to rapid increases in genetic variants
- Over 90% of the “virus” content of the P2 stock and 100% of the P3 stock (multiplicity of infection (MOI) ranging from 1.0E−02 to 1.0E−03) contained a 24-nucleotide in-frame deletion in the spike region resulting in loss of 8 amino acids including the FCS
- Virologists have become concerned that “SARS-CoV-2” isolates being propagated and studied around the globe were not accurately representing the “virus” circulating in humans
- The levels of FCS variants rose rapidly in Vero E6 cells irrespective of their source
- There is a risk that during the sequential passage of this “virus” for working stock generation, deletions may arise in critical virulence components of the “virus”
- They ultimately reccomend limiting of the number of passages in cell culture
- There are acceptable (<1%) variant thresholds and they recommend the inclusion of evaluations of minor variations based on consensus sequence (seemingly confirming there are always variations present)
- Deletions and mutations in the “SARS-CoV-2” FCS became so frequently observed in passages 4 and 5 that they dominated the reads taken from workings stocks by up to 99%
- Their data suggests that even the same passage level of “virus” can exhibit entirely different genetic characteristics
- They state that that investigators need to confirm the genetic sequence after propagation rather than relying on the sequence of the seed stock
- However. despite the publication of these articles, there is continued production and dissemination of stocks of “virus” that are compromised in this manner
As can be seen from the highlights from this first study, the culture conditions greatly influence the end results of the cell culture experiments. Serial propagation leads to dramatic increases in “variants” to the point that Virologists are concerned these stocks no longer resemble the circulating “virus.” They even advise researchers to always sequence their stock after propagation rather than relying on the original sequence due to the mutations which occur during the culturing process. However, not a single genome from any of these cultures ever matches 100%. They even seem to suggest that there is an acceptable level of variation. Even though the issues outlined above are known, the researchers admit that they are frequently overlooked and that there is continued production of “viral” stocks in this manner.

This second article sheds even more light on this problem:
SARS-CoV-2 variants with mutations at the S1/S2 cleavage site are generated in vitro during propagation in TMPRSS2-deficient cells
“Notably, viruses with S gene mutations emerged rapidly and became the dominant SARS-CoV-2 variants in TMPRSS2-deficient cells including Vero cells. Our study demonstrated that the S protein polybasic cleavage motif is a critical factor underlying SARS-CoV-2 entry and cell tropism. As such, researchers should be alert to the possibility of de novo S gene mutations emerging in tissue-culture propagated virus strains.”
“SARS-CoV-2 uses its spike (S) protein to enter target cells. Unlike other similar coronaviruses, the nascent S protein has a polybasic cleavage motif and is cleaved by the host protease. We have identified SARS-CoV-2 variants with mutations at the cleavage motif of S protein (S gene mutants) which undergo inefficient proteolytic cleavage, generate smaller plaques, and infect fewer cell lines. Notably, S gene mutants emerged rapidly through SARS-CoV-2 propagation in Vero cells. Since Vero cells are commonly used for SARS-CoV-2 propagation, it is a very real possibility that researchers have performed experiments, screened antivirals, and developed vaccines using SARS-CoV-2 S gene mutants without realizing.”
“In this study, we isolated S gene mutants from SARS-CoV-2 WK-521, a strain isolated from a clinical case in Japan [17], via serial passage in Vero cells. Other studies have reported viruses with S gene mutations, including amino acid deletions and substitution at the S1/S2 cleavage site from clinical isolates in Australia [21], China [20,22], England [23], and the USA [24] that emerged during cultivation in Vero cells or in its derivative, Vero/hSLAM, which are cells that do not express TMPRSS2. Although these studies demonstrated the spontaneous mutations in the S1/S2 cleavage site during the in vitro propagation of SARS-CoV-2, the virological properties of these mutants requires further investigation. This study showed a difference in cell tropism and entry pathway between SARS-CoV-2 WT and S gene mutants. We also demonstrated that the SARS-CoV-2 S gene mutants emerged within a few passages and became the dominant SARS-CoV-2 variants in TMPRSS2-deficient cells.”
“The spontaneous mutations in the S gene that lead to a loss of sensitivity to protease have been identified during the passage of cultured cells and this phenomenon is considered an adaptation of coronaviruses, such as human coronavirus OC43 and feline coronavirus UCD, to cell culture [18,38]. Our deep sequencing analysis revealed that S gene mutants emerged at P1 and rapidly became the dominant variant within the virus populations that emerged from Vero cell passage. Our findings indicate that replication of SARS-CoV-2 in TMPRSS2-deficient Vero cells results in the selection of S gene mutants; as such, passage in this cell line is technically inappropriate, as it becomes difficult to maintain SARS-CoV-2 with the S1/S2 cleavage site in its intact form.”
“At this time, many studies are conducted using SARS-CoV-2 propagated in Vero cells. Considering the very real possibility that these virus stocks will accumulate S gene mutations, researchers must pay careful attention to the passage history of any working stocks of SARS-CoV-2. Moreover, we must be very objective when interpreting the results from studies using Vero-passaged virus, especially those focused on S protein cleavage, virus entry and on cell tropism of SARS-CoV-2.”
https://journals.plos.org/plospathogens/article?id=10.1371/journal.ppat.1009233
In Summary (Part 2):
- Researchers should be alert to the possibility of de novo S gene mutations emerging in tissue-culture propagated “virus” strains
- S gene mutants emerged rapidly through “SARS-CoV-2” propagation in Vero cells
- Since Vero cells are commonly used for “SARS-CoV-2” propagation, it is a very real possibility that researchers have performed experiments, screened antivirals, and developed vaccines using “SARS-CoV-2” S gene mutants without realizing it
- The researchers state that they demonstrated that the “SARS-CoV-2” S gene mutants emerged within a few passages and became the dominant “SARS-CoV-2” variants in TMPRSS2-deficient cells
- They state that their findings indicate that replication of “SARS-CoV-2” in TMPRSS2-deficient Vero cells results in the selection of S gene mutants; as such, passage in this cell line is technically inappropriate, as it becomes difficult to maintain “SARS-CoV-2” with the S1/S2 cleavage site in its intact form
- They conclude that researchers must be very objective when interpreting the results from studies using Vero-passaged “virus“
From this study, the researchers claim that not only does passaging in Vero cells lead to mutations, it does so in the infamous spike (S) protein. And just as the previous paper stated, they claim that this is so common that researchers are most likely using these mutants in their research without ever realizing it. They also warn that results from Vero-passaged studies must be interpreted very objectively.
If one looks at this critically and logically, it is clear that the cell culture conditions greatly influence the results of the experiments and data generated from them. Cell cultures are unnatural mixtures of human/animal DNA, numerous chemicals/antibiotics, nutrients, etc. that are carried out in laboratory conditions that have no relation to reality whatsoever. The variations and mutations will remain as no two culture conditions are ever the exact same and thus the results will always be different. There can be no claim that what ultimately results from the cell culture process is the same as what went in at the beginning. In fact, the evidence points to the fact that this is never the case.


2 Responses
carolyn gutman dey
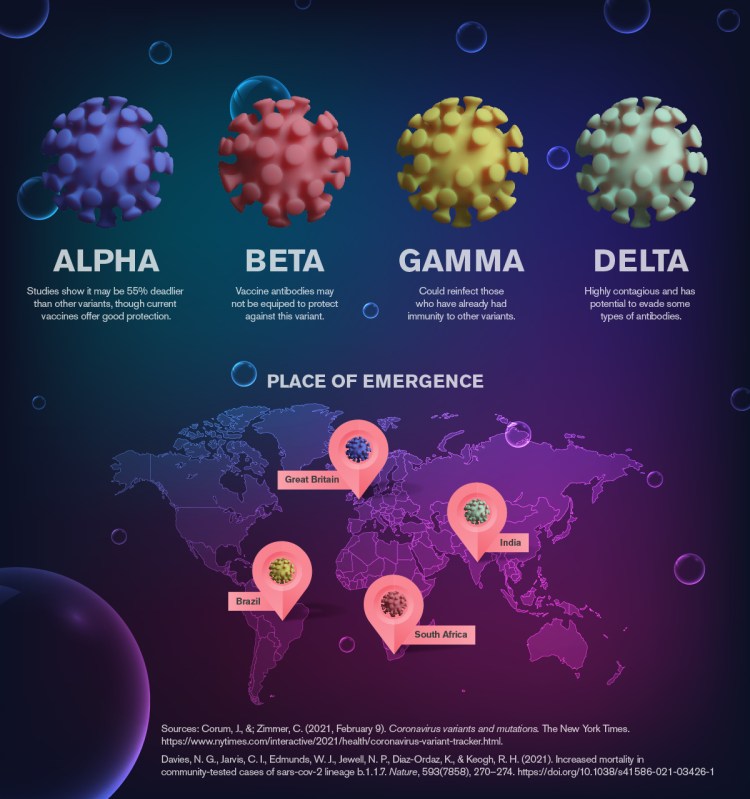
wow that image at the top is interesting….. the globe and the virus, what a pair!
Misto1481
Of course! They have to keep their fictional narratives together and supporting each other in the collective consciousness. 😉